BU Scientists Advance Efforts to Predict the Behavior of Bacterial Colonies
Using COMETS simulations, researchers showed how metabolism, nutrient use, and physical growth shape colony structure and genetic diversity

Microbes — tiny organisms invisible to the naked eye — rarely live in isolation. They form dynamic communities that shape ecosystems in our bodies, soils, oceans, and industries. Despite their microscopic size, these communities wield an outsized influence on human health, agriculture, and sectors ranging from biofuels to pharmaceuticals.
Both beneficial and harmful, microbes hold vast potential. Featured on the cover of the August issue of Cell Systems, a new Boston University study uncovers how microbial colonies grow and organize, advancing efforts to predict their behavior. By using computer simulations, the study shows that bacterial growth is not random but emerges from the interplay of metabolism, nutrient flow, and physical dynamics.

Building on their COMETS (Computation Of Microbial Ecosystems in Time and Space) framework, BU Professors Ilija Dukovski, Kirill Korolev and Daniel Segrè with collaborators created a new tool to simulate how microbial communities develop under different conditions. They modeled E. coli colonies by combining detailed metabolic models with the physical processes that drive colony expansion and organization. This approach captured both the metabolic activity of individual bacteria and the spreading behavior of the colony as a whole. The work shows how COMETS bridges theory and experiment, simulating microbial interactions and ecosystem changes beyond traditional models.

Study Findings
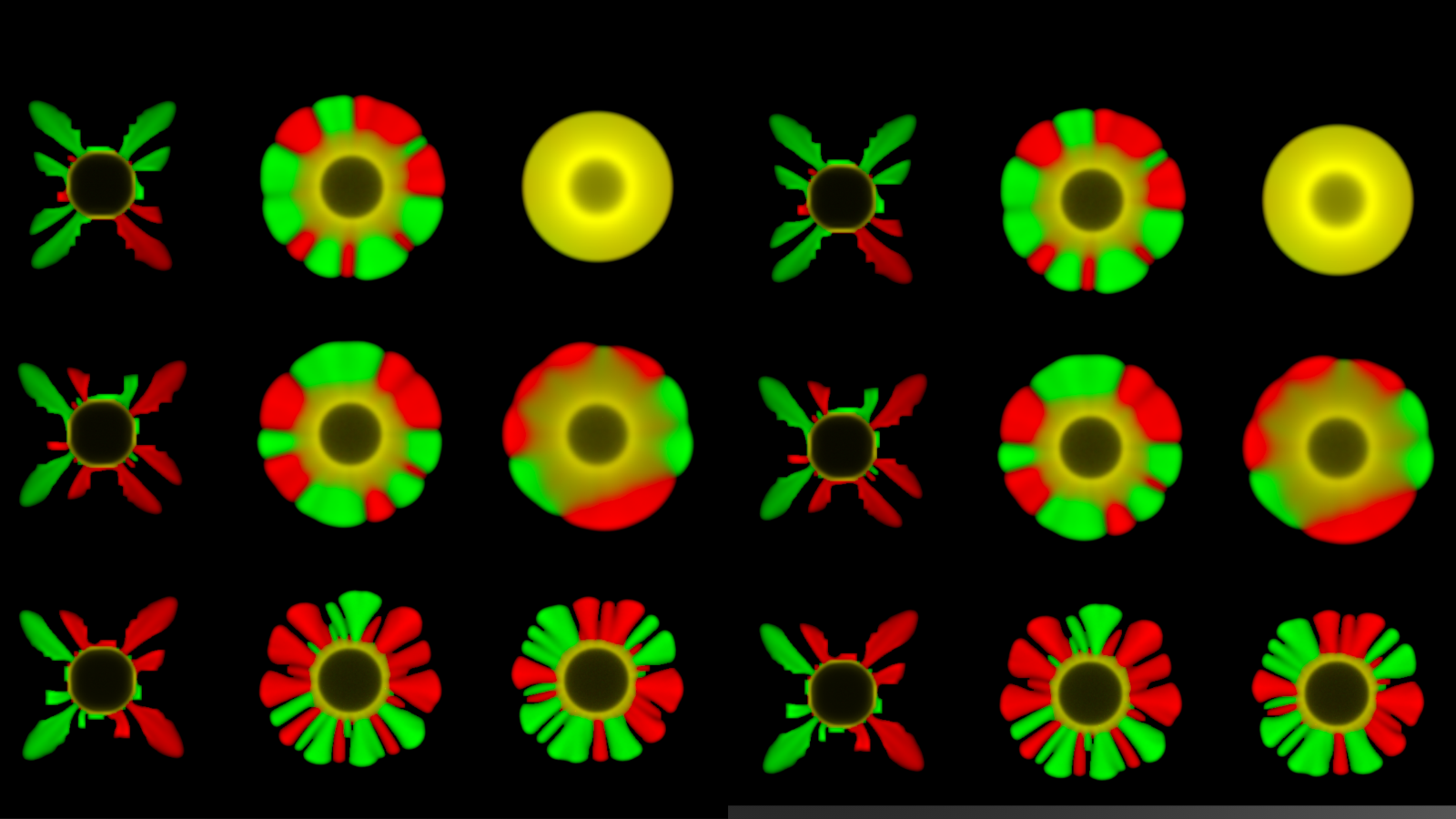
The study revealed that colony development depends not only on nutrient availability but also on the physical spreading of biomass — the way cells grow and expand as a colony over time — and the metabolic choices cells make.
The team also found that colony shape can affect genetic diversity. Branched colonies may isolate strains and preserve unique traits, while smoother colonies allow cells to mix more freely. Another striking observation was the formation of “metabolic rings”: dense, ring-shaped bands of growth resembling coffee-stain patterns, but caused by nutrient diffusion and metabolism rather than evaporation.

Overall, the simulations successfully reproduced lab-observed colony shapes, growth patterns, and the gradual decline of genetic diversity over time.
“These insights offer a new understanding of how bacteria maintain diversity and adapt to their surroundings,” says Daniel Segrè, corresponding study author; professor of biology, and of computing and data science; and Hariri Institute faculty affiliate at Boston University. “By uncovering these patterns, researchers can better predict how microbial communities respond to environmental changes, with applications in medicine, ecology, and biotechnology.”
Previous studies have explored either metabolism or physical growth, but none had combined both to capture the full picture of colony formation and organization.

“Our COMETS framework opens new ways to study microbial communities,” adds co-corresponding author Kirill S. Korolev, associate professor of physics and bioinformatics; and Hariri Institute faculty affiliate at Boston University. “We can now explore everything from how bacteria colonize plant roots to how blocking specific metabolic pathways alters growth, and even begin to predict how shifts in nutrients or metabolism influence entire communities.”
Looking ahead, the researchers plan to expand COMETS to study more complex microbial communities, including multiple species and intercellular interactions. Their work could provide insights into how bacteria colonize medical devices, plant roots, and soil, and could inform new strategies in biotechnology and medicine.
About COMETS
COMETS is a publicly available software that is developed and maintained in the Segrè Lab at Boston University. Its most recent major release was the outcome of a collaboration with Yale University and the University of Minnesota. A 2017 Hariri award was instrumental in early development of COMETS leading to this study. More information, including the COMETS user manual, how it can be downloaded and installed are found at the COMETS website: http://runcomets.org.
Paper citation: Ilija Dukovski, Lauren Golden, Jing Zhang, Melisa Osborne, Daniel Segrè, Kirill S. Korolev. Biophysical metabolic modeling of complex bacterial colony morphology. Cell Systems, Volume 16, Issue 8, 101352. https://doi.org/10.1016/j.cels.2025.101352.
This work was supported by the U.S. Department of Energy, Office of Science, Biological & Environmental Research (m-CAFEs, DE-AC02-05CH11231 to LBNL); the National Science Foundation (NSF-OCE-BSF 1635070; NSF Center for Chemical Currencies of a Microbial Planet); the Human Frontiers Science Program (RGP0060/2021); the NIH National Cancer Institute (1R21CA279630-01); the NIH National Institute of General Medical Sciences (1R01GM138530-01), and the Boston University Biological Design Center Kilachand Multicellular Design Program.