Research
It is becoming increasingly clear that complex genetic diseases manifest differently in each
person, tissue, and even cell, and that many of these differences are driven by subtle changes
in gene regulation. Vast amounts of multi-modal data are therefore needed to understand how
these diseases function and progress. As the scale and complexity of data available continues
to grow, cutting-edge methods at the intersection of computer science and genomics are
necessary for extracting biological meaning and making actionable predictions.
Network-based algorithms, which encode information as a network constructed from nodes and
edges representing the underlying data, are naturally suited for large-scale data integration.
Edges in a network can intuitively represent a wide array of relationships including between
individuals, cells, proteins, and genetic regions across multiple datasets.
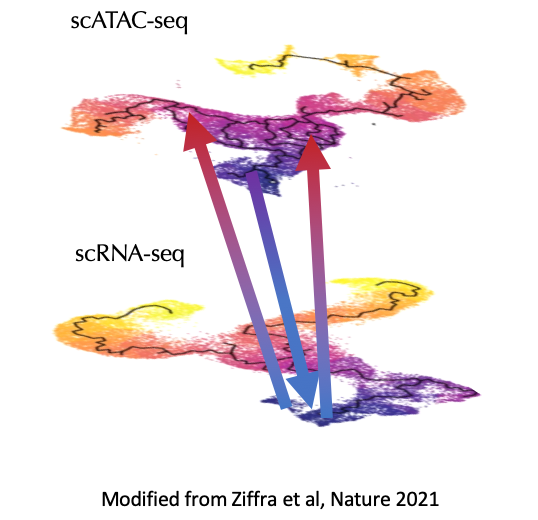
Continuous cell state analysis
Cell types are not discrete but rather are a complex mixture of states that vary continuously both across developmental time and with disease progression. To better understand changes in cell state, we are building network-based models for directed traversal through multi-modal data.
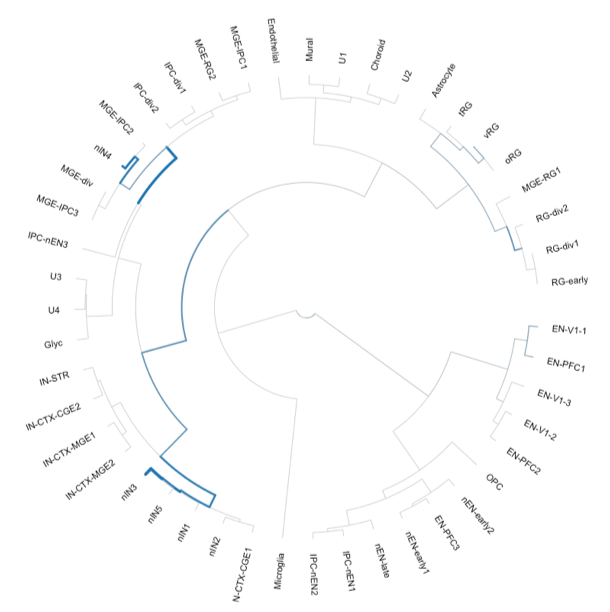
Interpretation of genetic variants
Complex diseases are caused by a heterogeneous and hard to interpret collection of genetic variants throughout the noncoding genome. We build tools to identify variants associated with changes in cell state by integrating variants with other genomic data.
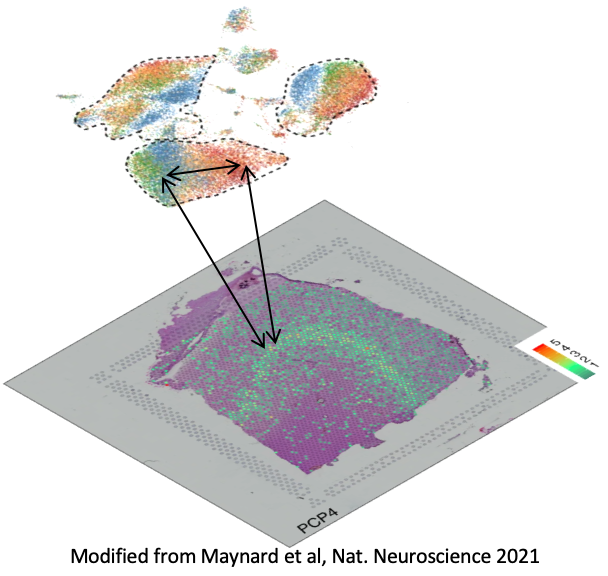
Models to integrate spatial data
Adding a spatial dimension to the already complex landscape of genetic regulation requires models that are capable of simultaneously considering physical distance and biological similarity. Networks we build based on these data allow for concrete determinations of how, where, and in which cells regulation is taking place.
User-friendly software
Encapsulating models that assist in the interpretation of data in user-friendly software is crucial for simplifying and increasing access to cutting-edge methods. We develop user-friendly software to close the gap between large-scale data collection and targeted experiments.