Overview
Hierarchical N-body algorithms
There are two common variants of hierarchical N-body algorithms: treecodes and FMMs. Both use tree data structures to cluster ‘source particles’ into a hierarchy of cells. See Figure 1.
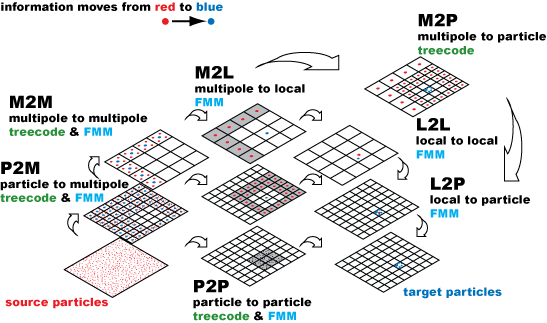
Multipole expansions are used to represent the effect of a cluster of particles, and are built for parent cells by an easy transformation (M2M). Local expansions are used to evaluate the effect of multipole expansions on many target points locally. The multipole-to-local (M2L) transformation is used only on the FMM, while treecodes compute the effect of multipole expansions directly on target points, with O(N log N) scaling for N particles. The FMM has the ideal O(N) scaling.
Parallel, multi-GPU, auto-tuning
We developed a new treecode-FMM hybrid algorithm with auto-tuning capabilities, that is O(N) and chooses the most efficient type of interaction. It is highly parallel and GPU-capable. For some details about the hybrid algorithm, see the Features of ExaFMM.
Parallel performance has been studied in both strong and weak scaling. On multi-core systemas (Kraken supercomputer), strong scaling with 108 particles on 2048 processes achieved:
- 93% parallel efficiency for the non-SIMD code, and
- 54% for the SIMD-optimized version (which is 2x faster).
Weak scaling with 106 particles per node achieved 72% efficiency on 32,768 processes. These results are detailed in the following publication:
“A tuned and scalable fast multipole method as a preeminent algorithm for exascale systems”, Rio Yokota and Lorena A Barba, Int. J. High-perf. Comput. Appl. (accepted 2011) Preprint arXiv:1106.2176
Summary of features
- Inter-node parallelism with MPI
- Intra-node multithreading with OpenMP
- GPU-enabled using CUDA
- Auto-tuning
- Released under the MIT License