Optimization and Learning Strategies for Protein Docking
Pirooz Vakili, Ioannis Paschalidis, Dima Kozakov, and Sandor Vajda
Supported by:
NIH R01 GM135930 “Optimization and Learning Strategies for Protein Docking” (PI: Pirooz Vakili)
NIH R35 GM118078 “Analysis and Prediction of Molecular Interactions” (PI: Sandor Vajda)
NSF DBI 1759472 “Collaborative Research: ABI Development: The next stage in protein‐protein docking” (PI: Sandor Vajda)
NIH R21 GM127952 “Fast Energy Evaluation For Multi-Protein Systems” (PI: Dima Kozakov)
Aim 1: Manifold Optimization. We have introduced a novel formulation of rigid body transformations that corresponds to the (Lie) group SO(3) × R3, i.e., the direct product of the (Lie) groups SO(3) and R3. 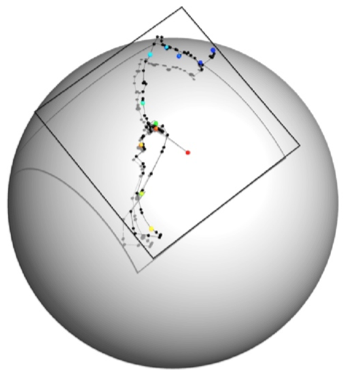 In our formulation, we select an initial center of rotation of the ligand p ∈ R3 (e.g., the center of mass of the interface between the ligand and the receptor). Given this choice, the rigid body transformation we associate with a rotation matrix R and translation y transforms a point q ∈ R3 on the ligand as q → R(q − p) + p + y. In this case, the rigid body optimization problem can be naturally defined as an optimization on SO(3)×R3. Efficient optimization algorithms for the component manifolds SO(3) and R3 exist and can be used to define efficient optimization algorithms for the product manifold SO(3)×R3. We have shown that this novel representation better captures the motion of the ligand with respect to the receptor and provides valuable flexibilities in the process of optimization. One of the main tasks of the project in the context of optimization is to use the new representation of rigid motion as well as the new Riemannian metric as the basis for our various manifold optimization algorithms. In what follows, we detail some of the specific components of this effort. We plan to fully extend our novel rigid-body representation to include a range of flexibilities. More specifically, we will integrate the 6D rotational/translational rigid moves with internal rotations around rotatable bonds within each molecule of the complex. We can represent these flexibilities using the internal coordinates of the ligand and receptor, and combine the internal coordinate representation with our manifold representation of the rigid moves. While the manifold parametrization defines a search space of minimal dimension for two arbitrary rigid bodies, proteins exhibit a lot of special structure enabling further dimensionality reduction in the vicinity of the native state.
In our formulation, we select an initial center of rotation of the ligand p ∈ R3 (e.g., the center of mass of the interface between the ligand and the receptor). Given this choice, the rigid body transformation we associate with a rotation matrix R and translation y transforms a point q ∈ R3 on the ligand as q → R(q − p) + p + y. In this case, the rigid body optimization problem can be naturally defined as an optimization on SO(3)×R3. Efficient optimization algorithms for the component manifolds SO(3) and R3 exist and can be used to define efficient optimization algorithms for the product manifold SO(3)×R3. We have shown that this novel representation better captures the motion of the ligand with respect to the receptor and provides valuable flexibilities in the process of optimization. One of the main tasks of the project in the context of optimization is to use the new representation of rigid motion as well as the new Riemannian metric as the basis for our various manifold optimization algorithms. In what follows, we detail some of the specific components of this effort. We plan to fully extend our novel rigid-body representation to include a range of flexibilities. More specifically, we will integrate the 6D rotational/translational rigid moves with internal rotations around rotatable bonds within each molecule of the complex. We can represent these flexibilities using the internal coordinates of the ligand and receptor, and combine the internal coordinate representation with our manifold representation of the rigid moves. While the manifold parametrization defines a search space of minimal dimension for two arbitrary rigid bodies, proteins exhibit a lot of special structure enabling further dimensionality reduction in the vicinity of the native state.
Aim 2: Development of a Medium Space-Scale Refinement Method. Here, we focus on optimization in the medium space-scale. We assume that initial global sampling in the entire conformational space has been performed by FFT-based methods such as PIPER implemented in the protein docking server ClusPro. These conformations are then sorted by their energy values, and the top few thousands with the lowest energy are retained for further processing, typically referred to as refinement. We have developed a stochastic global optimization method called Semi-Definite programming-based Underestimation (SDU), which improved upon earlier work. SDU works within a conformational cluster retained from FFT-based rigid docking. The key idea is to exploit the funnel-like structure of the energy function, pointed our earlier in the proposal, by first estimating the funnel and then biasing sampling towards the funnel basin, thereby, drastically increasing the sampling efficiency. We have established a number of mathematical results that rigorously prove SDU convergence as the number of samples grows. Here we propose to take full advantage of the reduced dimensionality concept. In the restrictive subspace identified in the process of progressive dimensionality reduction, there is not much scope for optimization as the function resembles a very steep canyon. Hence, the sampling of points with a non-zero projection along the restrictive eigenvector coordinates results in severe steric overlaps and is inefficient. More specifically, having identified the permissive subspace by PCA, we plan to develop an algorithm that only samples in this subspace and simply uses the cluster average values for the coordinates in the restrictive subspace. We call such a method, the Subspace Semi-Definite programming-based Underestimation (SSDU). As SSDU iterates, it will sample a sequence of different permissive subspaces, essentially capturing some of the association dynamics and following the preferred association pathway guiding the two proteins towards the native state.
Aim 3: Employing Learning Strategies. We focus on using learning methods at the conclusion of the refinement stage. First, we apply ML to cluster selection and ranking. The starting point for the work is a refined ensemble of conformations produced with the methods we outlined earlier. The objective is to select a small number of conformations with the highest probability of being near-native. This task is very challenging due to the inaccuracy of the energy potentials used in docking. Part of the inaccuracy comes from the fact that these potentials do not incorporate metrics of entropy, in addition to many other approximations that are used in energy calculations. To address this issue, we will employ learning techniques to combine energy with many other features and use them in discrimination and ranking. In our preliminary work referred to above, we only used cluster-dependent features relating to energy distribution, cluster size, and density. We plan to add a variety of other relevant features, exploiting the fact that each cluster of conformations we have formed implicitly specifies a set of atoms in the interface between the interacting proteins. Properties of this interface can introduce additional information to be used in the course of cluster ranking. Motivated by template-based docking (TBD) methods, we also plan to add features that indicate good agreement with an existing template.
Publications:
Mirzaei H, Zarbafian S, Villar E, Mottarella S, Beglov D, Vajda S, Paschalidis ICh, Vakili P, Kozakov D. Energy Minimization on Manifolds for Docking Flexible Molecules. J Chem Theory Comput. 2015 Mar 10;11(3):1063-76.
Nan, F.; Moghadasi, M.; Vakili, P.; Vajda, S.; Kozakov, D.; Paschalidis, I. C. A Subspace Semi-Definite programming-based Underestimation (SSDU) method for stochastic global optimization in protein docking. Proc IEEE Conf Decis Control 2014, 4623-4628, doi:10.1109/CDC.2014.7040111
Vakili P, Mirzaei H, Zarbafian S, Paschalidis IC, Kozakov D, Vajda S. Optimization on the space of rigid and flexible motions: an alternative manifold optimization approach. Proc IEEE Conf Decis Control. 2014 Dec;2014:5825-5830.
Kozakov, D.; Li, K.; Hall, D.R.; Beglov, D.; Zheng, J.; Vakili, P.; Schueler-Furman, O.; Paschalidis, I.; Clore, G.M.; Vajda, S. Encounter complexes and dimensionality reduction in protein-protein association. Elife 2014, 3, e01370, doi:10.7554/eLife.01370.
Mirzaei, H.; Villar, E.; Mottarella, S.; Beglov, D.; Paschalidis, I.C.; Vajda, S.; Kozakov, D.; Vakili, P. Flexible Refinement of Protein-Ligand Docking on Manifolds. Proc IEEE Conf Decis Control 2013, 1392-1397, doi:10.1109/CDC.2013.6760077.
Moghadasi M, Kozakov D, Vakili P, Vajda S, Paschalidis IC. A New Distributed Algorithm for Side-Chain Positioning in the Process of Protein Docking. Proc IEEE Conf Decis Control. 2013:739-744.
Mirzaei, H.; Beglov, D.; Paschalidis, I.C.; Vajda, S.; Vakili, P.; Kozakov, D. Rigid body energy minimization on manifolds for molecular docking. Journal of chemical theory and computation 2012, 8, 4374-4380, doi:10.1021/ct300272j.
Mirzaei H, Kozakov D, Beglov D, Paschalidis IC, Vajda S, Vakili P. A New Approach to Rigid Body Minimization with Application to Molecular Docking. Proc IEEE Conf Decis Control. 2012 Dec:2983-2988.
Moghadasi M, Kozakov D, Mamonov AB, Vakili P, Vajda S, Paschalidis IC. A Message Passing Approach to Side Chain Positioning with Applications in Protein Docking. Proc IEEE Conf Decis Control. 2012:2310-2315.
Kozakov D, Hall DR, Beglov D, Brenke R, Comeau SR, Shen Y, Li K, Zheng J, Vakili P, Paschalidis ICh, Vajda S. Achieving reliability and high accuracy in automated protein docking: ClusPro, PIPER, SDU, and stability analysis in CAPRI rounds 13-19. Proteins. 2010 Nov 15;78(15):3124-30. doi: 10.1002/prot.22835.
Shen Y, Paschalidis ICh, Vakili P, Vajda S. Protein docking by the underestimation of free energy funnels in the space of encounter complexes. PLoS Comput Biol. 2008 Oct;4(10):e1000191. doi: 10.1371/journal.pcbi.1000191.
Paschalidis ICh, Shen Y, Vakili P, Vajda S. Protein-protein docking with reduced potentials by exploiting multi-dimensional energy funnels. Conf Proc IEEE Eng Med Biol Soc. 2006;1:5330-3.
Paschalidis IC, Shen Y, Vakili P, Vajda S. SDU: A Semidefinite Programming-Based Underestimation Method for Stochastic Global Optimization in Protein Docking. IEEE Trans Automat Contr. 2007 Apr 1;52(4):664-676.