News
Congratulations to Soyoung Bae on winning a Stanford PULSE Scholarship
Soyoung Bae, a third-year graduate student in the Molecular Biology, Cell Biology, and Biochemistry (MCBB) PhD Degree program, has been awarded the prestigious Stanford PULSE Scholarship for the Ultra-Fast X-ray Summer School (UXSS) in June 2024. The UXSS program, hosted by Stanford's PULSE Institute at the Stanford Linear Accelerator Center (SLAC) National Accelerator Laboratory, is renowned for its cutting-edge training in ultrafast (atto–femto second) X-ray science, and Soyoung’s selection is a testament to her exceptional research and potential in the field of structural biology. The award covered her lodging at the Stanford workshop, allowing her to participate in this highly esteemed program.
Soyoung’s research in the laboratory of Biology Professor Dean Tolan, where she employs sophisticated techniques such as X-ray crystallography and surface plasmon resonance, aligns perfectly with the innovative focus of UXSS. Her participation in this program will undoubtedly enhance her skills and knowledge, further advancing her impactful research on enzyme structure and function.
Beyond BU for the Summer: Alexandra Lion Completes Intensive Embryology Course
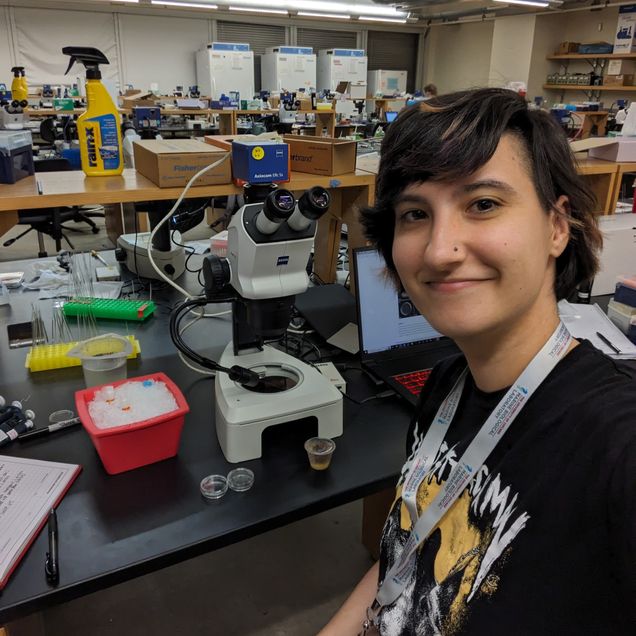

Alexandra Lion is a PhD candidate in the Bradham Lab, which studies pattern formation during embryonic development using the larval skeleton of the sea urchin Lytechinus variegatus as a model. Her projects focus on the role of the DEAD-box RNA helicase DDX6 in temporal regulation of development, and on the effect of per- and polyfluoryl alkyl substances (PFAS) on embryonic development and patterning.
Alexandra recently spent part of her summer attending the Marine Biological Laboratory course, Embryology: Concepts and Techniques in Modern Developmental Biology. The Embryology Course is an intensive six-week laboratory and lecture course for advanced graduate students, postdoctoral fellows, and more senior researchers who seek a broad and balanced view of modern issues in developmental biology.
Alexandra describes her experience below:
I applied to the Embryology course at the Marine Biological Laboratory because I wanted to expand my knowledge and connections within the field of developmental biology, as well as get more hands-on experience with a greater number of model organisms. Through this 6-week course, I was exposed to a wide range of well-established models such as Drosophila, zebrafish and C.elegans, as well as emerging model systems like acoels, cephalopods and ctenophores. I was able to greatly expand my microscopy experience since we were provided dozens of microscopes from vendors including Nikon, Leica, Andor and Zeiss. Having access to such a wide range of tools and organisms allowed me to try experiments that were outside of my prior expertise, such as lineage tracing, photo conversion, laser ablation, tissue grafting and more. This course really reinvigorated my sense of curiosity and opened my eyes to just how many unanswered questions there still are to probe. It helped that the course directors and faculty were enthusiastic about our ideas, and encouraged us to try any experiment we wanted to no matter how out-of-the-box. The faculty and course assistants frequently stayed in the lab with us students late into the night to help troubleshoot our experiments, and really supported all of our efforts.
The course introduced me to many people in the field, from other graduate students and postdocs to faculty who have been doing research for decades. Getting to work with the faculty in close quarters for up to a week at a time was incredibly valuable; you really get a feel for how the faculty are to work with, and how they might be as mentors. I also made a lot of really incredible friends through my time at Embryology; spending 6 weeks in the lab together, sharing all of our triumphs and setbacks at the bench, but also doing non-science things like practicing for the Embryology vs. Physiology softball game together really bonded us all. I’m grateful to have these people as both my friends and colleagues, and am excited to reunite at future conferences and events.
My attendance at the course was funded in part by a Max M. Burger Endowed Scholarship in Embryology and by the Society for Developmental Biology. Thank you also to the Biology Department and MCBB Program for their support, and to my advisor Cynthia Bradham for her encouragement and support of my attendance.
Katie Breuckman Accepted into NSF Graduate Research Traineeship Program in Biological Feedback Control

Katie Breuckman of the Hao and Khalil Labs was accepted as a trainee in the NSF Graduate Research Traineeship Program in Biological Feedback Control for the 2024-2025 academic year. This program is an NSF funded research traineeship combining the study of the engineering principles in feedback control with investigations of how biological systems self-regulate, adapt, heal and evolve. The NSF Research Traineeship (NRT) program seeks to provide effective training of STEM graduate students in high priority interdisciplinary or convergent research areas. Selected students participate in an industry internship, professional development events, and a co-mentored dissertation project in a breadth of topic areas including molecular level control algorithms, microbial feedback control systems, control of tissues homeostasis and regeneration, biohybrid and bioinspired engineered systems.
Katie is developing yeast-based selection methods for improving protease and nanobody specificity toward therapeutic targets. She aims to integrate laboratory evolution and scalable screening methods to rapidly, and cost-effectively produce bespoke therapeutics. Furthermore, through the NRT in Biological Feedback Control, she plans to develop mathematical models to predict evolutionary trajectories and computationally design improved starting points for further protein evolution. Through this work, Katie aims to enhance our grasp of feedback-based control for navigating protein fitness landscapes.
Congratulations, Katie!
Anna Berenson Receives 2024 Belamarich Award
 Dr. Anna Berenson of the Fuxman Bass Lab was selected as the winner of the Biology Department's 2024 Belamarich Award for her doctoral dissertation in Molecular Biology, Cell Biology & Biochemistry titled “Paired Yeast One-Hybrid Assays to Detect DNA-Binding Cooperativity and Antagonism Across Transcription Factors.” The selection committee was impressed by the quality of Anna’s work, its combination of methods development, observational studies, and computational analyses, and its potential for informing future research in the field of gene regulation. More information about her research is below.
Dr. Anna Berenson of the Fuxman Bass Lab was selected as the winner of the Biology Department's 2024 Belamarich Award for her doctoral dissertation in Molecular Biology, Cell Biology & Biochemistry titled “Paired Yeast One-Hybrid Assays to Detect DNA-Binding Cooperativity and Antagonism Across Transcription Factors.” The selection committee was impressed by the quality of Anna’s work, its combination of methods development, observational studies, and computational analyses, and its potential for informing future research in the field of gene regulation. More information about her research is below.
For her dissertation, Anna developed paired yeast one-hybrid (pY1H) assays to study interactions between pairs of transcription factor (TF) proteins and DNA regions of interest. In addition to identifying cooperative DNA binding of TF pairs, pY1H assays also revealed extensive DNA-binding antagonism between TFs, constituting a previously underappreciated mechanism to regulate TF-DNA binding. Anna further applied pY1H assays to study the role of TF-TF relationships in cytokine gene regulation, the effect of alternative TF isoform usage on these relationships, and the effect of viral proteins on human TF-DNA binding. This work contributes to our understanding of how TF-DNA interactions are specified and provides a useful method that can be applied to further elucidate TF-TF relationships and their role in transcriptional regulation.
Anna will be continuing her academic career as a postdoctoral researcher in the lab of Dr. Jef Boeke at the NYU Langone Health Institute for Systems Genetics.
Congratulations, Anna!
Michelle Teplensky Receives Beckman Young Investigator Award

Dr. Michelle Teplensky was recently named one of the 2024 Beckman Young Investigators. She and her team will receive $600,000 over four years to engineer versatile vaccine responses through nanomaterial design, with implications for the treatment of not only the flu but also other infectious diseases including the viruses that cause COVID-19, HIV, and more.
The Beckman Young Investigator (BYI) Program provides research support to the most promising young faculty members in the early stages of their academic careers in the chemical and life sciences, particularly to foster the invention of methods, instruments and materials that will open up new avenues of research in science.
Read a detailed announcement here.
Congratulations, Dr. Michelle Teplensky!
Jillian Ness Receives 2024 Marion R. Kramer Scholarship

Jillian Ness of the Wunderlich Lab is one of the recipients of the Biology Department's 2024 Marion R. Kramer Scholarship. Jillian and her team studied how enhancers work in development, focusing on redundant enhancers, or “shadow enhancers,” linked to developmental genes. These enhancers are remarkably abundant in animals and can compensate under conditions of stress to drive normal development.
She is exploring how these enhancers function, as well as how they are created and maintained in animal genomes. Her work involves creating simplified enhancer models in Drosophila and analyzing how they work. In parallel, she performs evolutionary studies on shadow enhancer sequences to understand genomic events from which the sequences originate. Ultimately, she aims to understand enhancer sequences to improve predictions of perturbations that lead to developmental disease in embryos.
Read the full announcement here.
Congratulations, Jillian!
Mandy Pinheiro Receives 2024 Belamarich Dissertation Writing Award

Mandy Pinheiro of the Naya Lab received the Biology Department's 2024 Belamarich Dissertation Writing Award. This award complements the Belamarich Award, and is given to support an outstanding PhD student through the dissertation writing stage.
Mandy’s research focuses on understanding how noncoding RNAs (ncRNAs) may be central to the coordination of metabolism and differentiation in skeletal muscle. The regulation of gene expression during cell state transitions is a complex and tightly controlled molecular process. One potential central regulator of myogenesis is the Dlk1-Dio3 ncRNA locus, the largest known mammalian cluster of ncRNAs. She used mouse skeletal muscle cells as a model system to investigate how this ncRNA cluster may coordinate metabolic and epigenetic changes that occur when proliferating myoblasts differentiate into mature myotubes.
Congratulations, Mandy!
Green Lab Publishes Patent for Detecting Nucleic Acids
Milad Babaei, MCBB PhD student of the Green Lab, and Dr. Zhaoqing Yan (GRS '23), along with their mentor, Dr. Alexander Green, are co-authors on a recently published patent. This patent, titled "Isothermal Nucleic Acid Detection Assays and Uses Thereof" by Green et al. (US Patent 2023/0313283, October 2, 2023), describes rapid methods for detecting nucleic acids, from viruses to circulating mRNAs, in low-cost portable formats.
Nucleic acids, such as DNA and RNA, play a crucial role in genetic information. "Isothermal" refers to processes that occur at a constant temperature. In the context of nucleic acid detection, isothermal assays are methods that allow the amplification and detection of nucleic acids at a constant temperature without the need for thermal cycling, as in PCR (Polymerase Chain Reaction).
The team is currently expanding its panel of detection by developing lateral flow strips that can spontaneously probe for multiple targets using multiple test lines. This capability is crucial for the future of detecting and analyzing nucleic acids, which are fundamental in various fields, including medical diagnostics, research, and biotechnology.
Congratulations, Dr. Alexander Green, Milad Babaei, and Dr. Zhaoqing Yan!
Michelle Teplensky Named AIChE’s 35 Under 35
 Dr. Michelle Teplensky was recently named 2023 AIChE's (American Institute of Chemical Engineers) 35 under 35. AIChE honors 35 chemical engineering professionals who have mad great contributions to the field in seven categories, bioengineering, chemicals and materials, education and outreach, energy and environment, innovation and entrepreneurship, leadership. and safety. Teplensky was recognized under the bioengineering category.
Dr. Michelle Teplensky was recently named 2023 AIChE's (American Institute of Chemical Engineers) 35 under 35. AIChE honors 35 chemical engineering professionals who have mad great contributions to the field in seven categories, bioengineering, chemicals and materials, education and outreach, energy and environment, innovation and entrepreneurship, leadership. and safety. Teplensky was recognized under the bioengineering category.
Teplensky and her research group focuses on engineering nanotechnology to control immunological cell connectivity, processing, and communication by design. In doing so, they elucidate fundamentals about cellular events and leverage this knowledge to develop improved therapeutics and vaccines that can impact the treatment of cancer and infectious disease. Their work is highly interdisciplinary, and bridges the fields of engineering, chemistry, nanotechnology, immunology, and biomaterials. By sitting at this interface, we are able to develop and incorporate synthetic nanoscale advances to control immunological activity and elucidate design rules that have widespread impacts on therapeutic development. She hopes that the students trained in her lab enter their careers feeling inspired.
Teplensky loves to shop at Stew Leonard’s, and she has become an avid fan of Boston Univ.’s hockey team. Her family has a tradition of buying and completing a puzzle on every vacation.
Congratulations, Michelle Teplensky!
Anna Berenson Receives First Place in the PhD on Tap 3MT Competition

Anna Berenson, an MCBB PhD candidate of the Fuxman Bass Lab, received joint first place in the 2023 PhD on Tap 3MT Competition. The competition took place as part of PhD on Tap, which is a full day research showcase sponsored by the Office of the Associate Provost for Graduate Affairs and the Graduate School of Arts and Sciences.
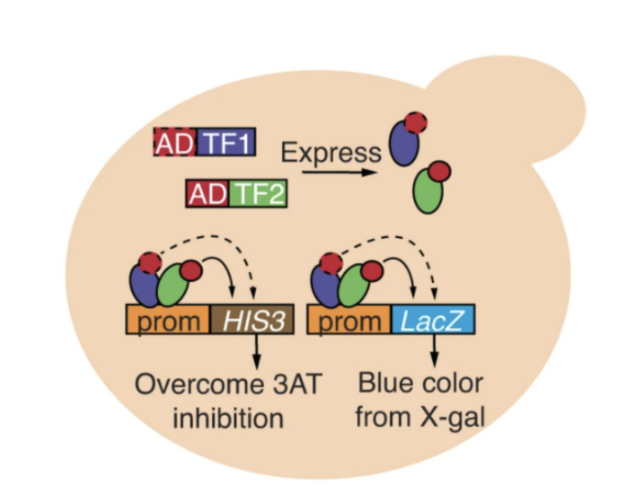
In a project led by Anna, the Fuxman Bass Lab developed paired yeast one-hybrid assays to study DNA binding patterns of transcription factor pairs. Their work shows that extensive cooperativity and antagonism between transcription factors greatly affect their binding to gene promoters, and that viral proteins can modify the binding profiles of human transcription factors. You can check out their publication here.
Congratulations, Anna!
